Welcome to the project page for TBTK
TBTK is a c++ library for solving general bilinear Hamiltonians. A flexible indexing scheme allows for general models to be setup with little effort. The model can either be solved using a DiagonalizationSolver, giving eigenstates and eigenvectors, or using a ChebyshevSolver to calculate Green's functions.
Note: The library is in beta stage and rapid changes to the code base can occur. It is therefore advised that a local copy of the library is created before updating it in case important projects are dependent on it.
Introductory material
Quick start
Presentation
Templates (see src/main.cpp in each template folder)
API
Example
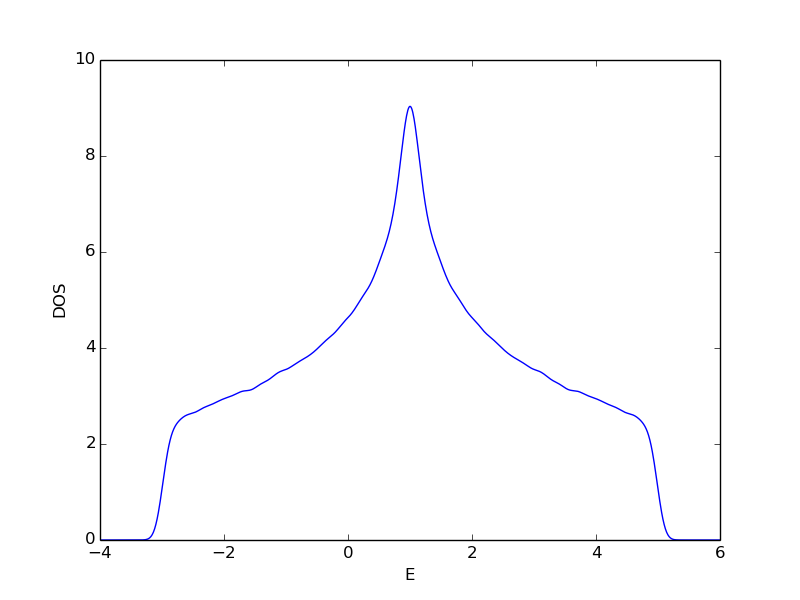
Setting up and diagonalizing a two dimensional tight-binding model with chemical potential mu and nearest neighbor hopping t. Eigenvalues and DOS are extracted and written to file.
//Create model and set up hopping parameters
Model model;
for(int x = 0; x < SIZE_X; x++){
for(int y = 0; y < SIZE_Y; y++){
for(int s = 0; s < 2; s++){
//Add hopping amplitudes corresponding to chemical potential
model.addHA(HoppingAmplitude(-mu, {x, y, s}, {x, y, s}));
//Add hopping amplitudes corresponding to t
if(x+1 < SIZE_X)
model.addHAAndHC(HoppingAmplitude(-t, {(x+1)%SIZE_X, y, s}, {x, y, s}));
if(y+1 < SIZE_Y)
model.addHAAndHC(HoppingAmplitude(-t, {x, (y+1)%SIZE_Y, s}, {x, y, s}));
}
}
}
//Construct model
model.construct();
//Setup and run DiagonalizationSolver
DiagonalizationSolver dSolver;
dSolver.setModel(&model);
dSolver.run();
//Set filename and remove any file already in the folder
FileWriter::setFileName("TBTKResults.h5");
FileWriter::clear();
//Create DPropertyExtractor
DPropertyExtractor pe(&dSolver);
//Extract eigenvalues and write these to file
Property::EigenValues *ev = pe.getEigenValues();
FileWriter::writeEigenValues(ev);
delete ev;
//Extract DOS and write to file
const double UPPER_LIMIT = 6.;
const double LOWER_LIMIT = -4.;
const int RESOLUTION = 1000;
Property::DOS *dos = pe.calculateDOS(UPPER_LIMIT, LOWER_LIMIT, RESOLUTION);
FileWriter::writeDOS(dos);
delete dos;
DOS calculated using SIZE_X=SIZE_Y=40 and plotted using Gaussian smoothing with sigma=0.1.
Installation
Currently the library can only be installed without modification if the computer has gcc 4.9 and nvcc 7.0 installed, and these can be loaded using "module load gcc/4.9" and "module load nvcc/7.0". If this is the case, type
git clone https://github.com/dafer45/TBTK
cd TBTK
source init_session.sh
./install.sh -CUDA
If CUDA is not available it is also possible to install the library without support for GPU by typing
git clone https://github.com/dafer45/TBTK
cd TBTK
source init_session.sh
./install.sh
If you are interested in installing the library on a different system, contact kristofer.bjornson@physics.uu.se for further instructions.
Update
To update to the latest version of the library, type
git pull
source init_session.sh
./update.sh -CUDA
To update the library without CUDA support, type
git pull
source init_session.sh
./update.sh
Initialize session
Each time a new session is started by opening a new shell, the library has to be initialized by entering the library folder and typing
source init_session.sh
Compile and run Templates
Enter the template folder, for example Templates/BasicDiagonalization, and type
make
./build/a.out
The results are now stored in the file TBTKResults.h5. These can further be plotted and saved in the folder figures by typing
./plot.sh
Note that plotting only is possible if python as well as the python libraries matplotlib, scipy, and h5py are installed on the computer.